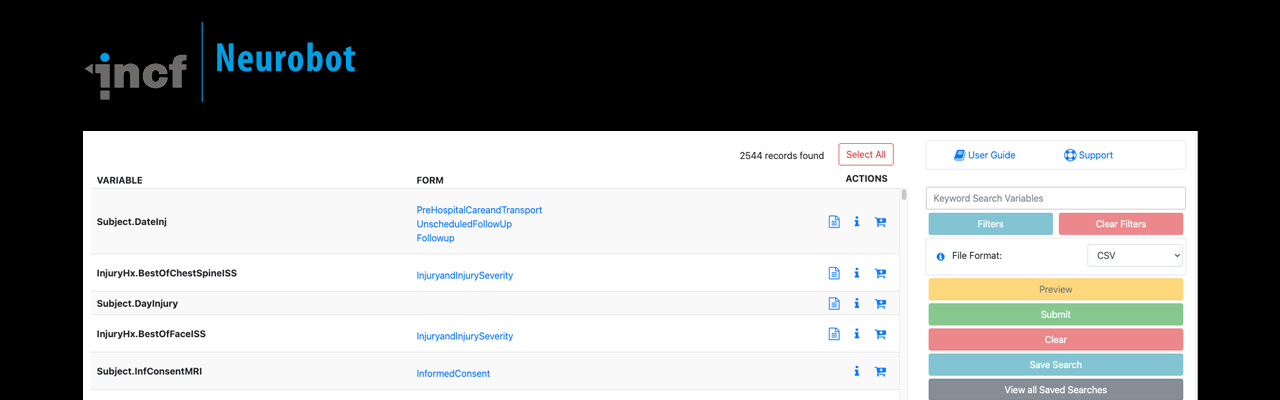
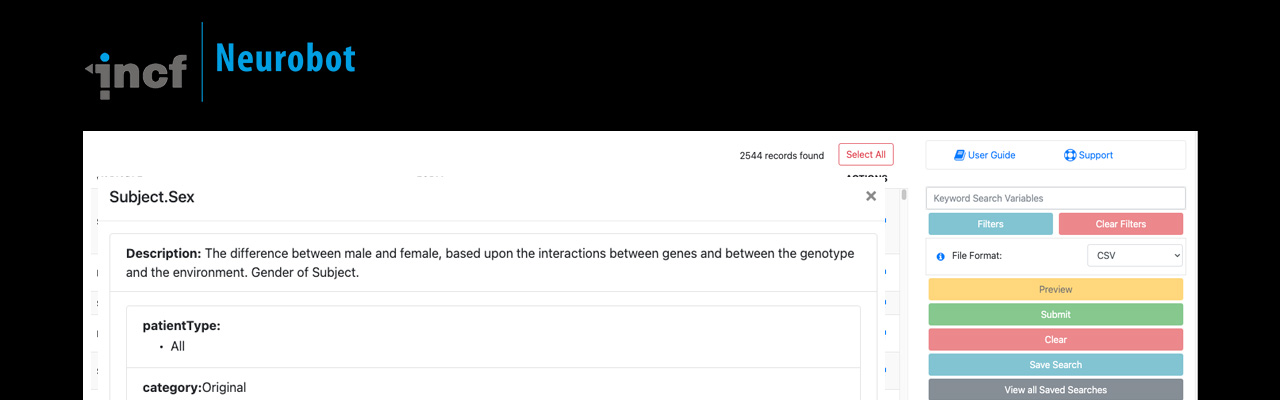
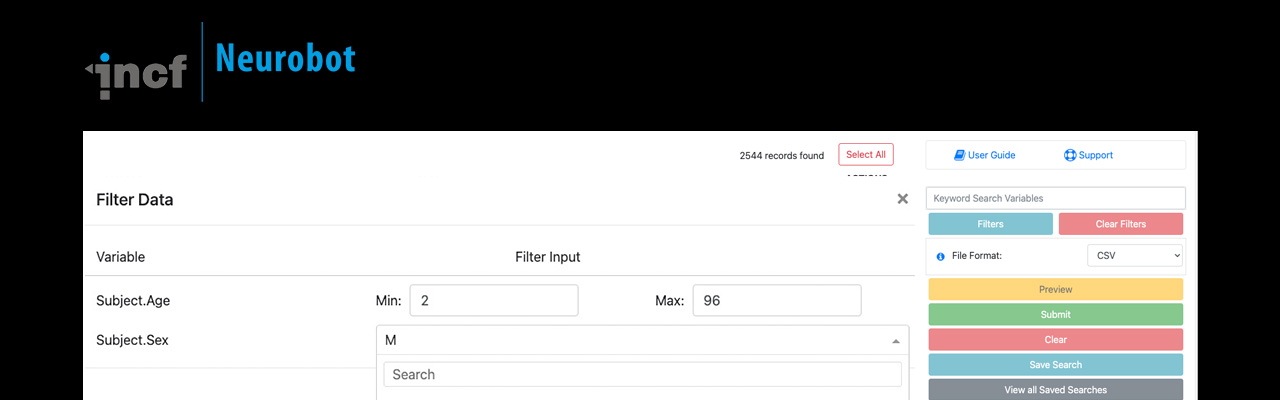
Neurobot is a web based application for simplifying data sharing and metadata management for research.
Most clinical data management tools are designed for efficient data acquisition and data processing; however, they often lack a usable data access interface. Neurobot, a lightweight data sharing application, was developed to provide a user-friendly data access interface that can be used for sharing a wide variety of versioned datasets.
The data model behind Neurobot has a scalable backend and has been optimised for faster queries on large datasets. By separating data publishing and sharing tools from the data management platform, Neurobot provides the flexibility required for clinical and non clinical studies. Check out the neurobot demo instance.
Neurobot for Study data managers, Researchers and Platform administrators
Why Neurobot?
Here is a short video describing the key features of Neurobot and how it can help with data sharing and metadata management for researchers.
Register for a Neurobot demo account today!
Key Features
Affiliated Studies
Use Case
A research lab has been saving the experiment values and results in csv files. For sharing the data with researchers. A shared folder is used where all researchers have access to. There has to be several knowledge transfer meetings to explain what the data elements are and how it can be accessed.
With Neurobot, the csv files can be uploaded by the data manager. After upload, the data manager can describe the data in the dictionary that is generated for each column in the csv. She can also give access to fellow researchers to update the dictionary. Data managers can also define different groups that can have access to the data.